Protein Evolution
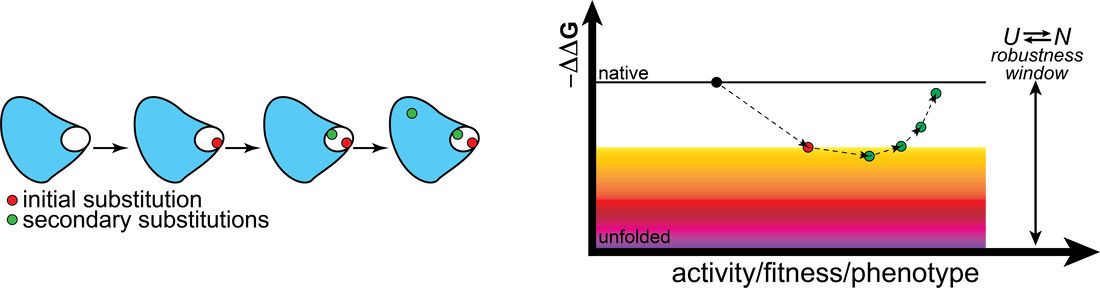
Our ability to understand protein evolution and the sequence-structure-function relationship as well as our success in enzyme engineering and the design of new functions hinges on understanding how evolutionary trajectories are shaped at the fundamental protein level. The activity-stability tradeoff model has transformed our view of enzyme evolution, however, the innovability-robustness diptych does not provide a complete molecular view of how the mechanism-based observables ΔΔG and kcat/KM, render specific evolutionary trajectories accessible. Instead, we combine molecular traits relevant to the statistical ensemble nature of protein structure, including low-populated states and motions at a wide range of timescales, with structural snapshots and in vivo readings to address questions like why secondary substitutions are fixed at specific positions within a large protein scaffold?
Chaperone assisted protein folding
We investigate the molecular mechanisms by which chaperones shape the conformational landscape of proteins in their nascent or mature states. The heat shock protein 90 (Hsp90), in cohort with other proteins termed co-chaperones, forms a machinery that acts at a late stage of protein folding. It confers fine structural changes to its substrates and induces their conformational maturation to a stable, functional and active state. Its “clients” comprise a variety of proteins involved in diverse cellular processes from signal transduction, to telomere maintenance and protein trafficking. Our long term goal is to understand the molecular mechanism by which different components of the Hsp90 machinery allow activation and afford stability of oncogenic kinases. Cdc37 (cell division cycle, 37) is a gain-of-specificity co-chaperone of the Hsp90 machinery, specializing in the recruitment of protein kinases. We hope that our mechanistic studies will ultimately lead to novel routes for a pharmacologic intervention in cancer.
Cdc37-mediated client kinase sorting
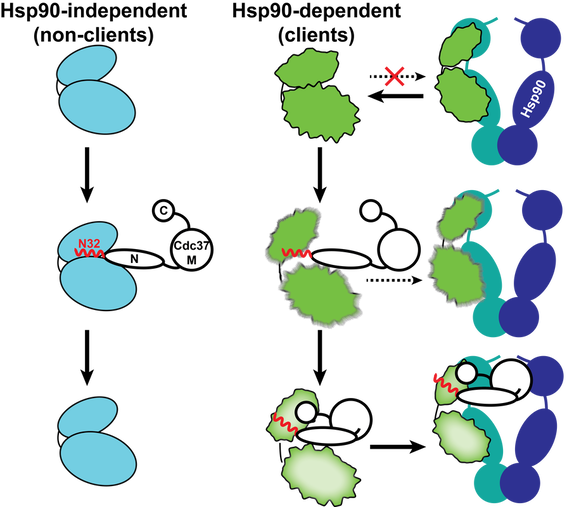
Cdc37 recognizes kinase specificity determinants in both client and nonclient kinases and thus acts as a general kinase scanning factor. However, the function of Cdc37 as a substrate recruiter cochaperone is not strictly that of an adaptor protein. Instead, it actively participates in making triage decisions as to which substrates require stable association with Hsp90 and it is therefore charged with the daunting task of kinase sorting within the client-nonclient continuum. Its sorting activity relies on its ability to challenge the conformational stability and locally unfolding only client kinases. This cochaperone-induced metastable conformational state acts as a “flag” for stable cochaperone association and thus Hsp90 dependence.
Keramisanou D, Aboalroub A, Zhang Z, Liu W, Marshall D, Diviney A, Larsen RW, Landgraf R, Gelis I.*
Molecular Mechanism of Protein Kinase Recognition and Sorting by the Hsp90 Kinome-Specific Cochaperone Cdc37
Molecular Cell, 62, 260-271 (2016)
Molecular Mechanism of Protein Kinase Recognition and Sorting by the Hsp90 Kinome-Specific Cochaperone Cdc37
Molecular Cell, 62, 260-271 (2016)
Phosphorylation-regulated kinase chaperoning
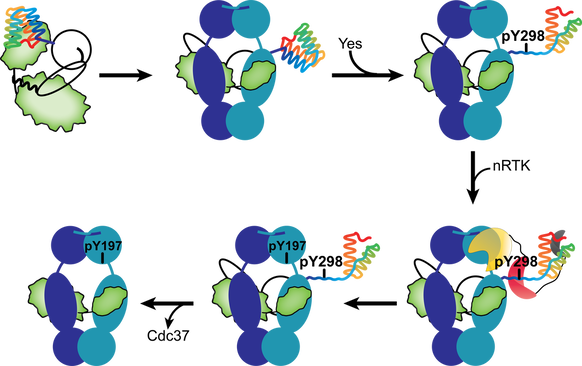
Phosphorylation alters protein function through versatile mechanisms that include allosteric changes, direct positive or negative modulation of protein-protein and protein-DNA interactions, availability of cofactor-binding sites, autoinhibition, disorder-to-order and order-to-disorder transitions. Phosphorylation of Cdc37 at Y298 results in partial unfolding of the C-terminal domain which unmasks a high-affinity SH2-binding phosphopeptide with broad specificity over multiple SH2 domains of nRTKs. Recruitment of Yes kinase via its SH2 (and SH3) domain increases its effective concertation at the complex, which promotes Hsp90 phosphorylation at Y197 (Hsp90α) and in turn results in Cdc37 disassembly. Therefore, by providing client class specificity, Hsp90 cochaperones, such as Cdc37, do not merely assist in client recruitment but also shape the post-translational modification landscape of Hsp90 in a client class-specific manner.
Bachman A.B., Keramisanou D., Xu W., Beebe K., Moses M.A., Vasantha Kumar M.V., Gray G., Noor R.E., van der Vaart A., Neckers L., Gelis I.*
Phosphorylation induced cochaperone unfolding promotes kinase recruitment and client class-specific Hsp90 phosphorylation
Nature Communications, 9, 265 (2018)
Phosphorylation induced cochaperone unfolding promotes kinase recruitment and client class-specific Hsp90 phosphorylation
Nature Communications, 9, 265 (2018)
Dynamic character of Hsp90 chaperoning
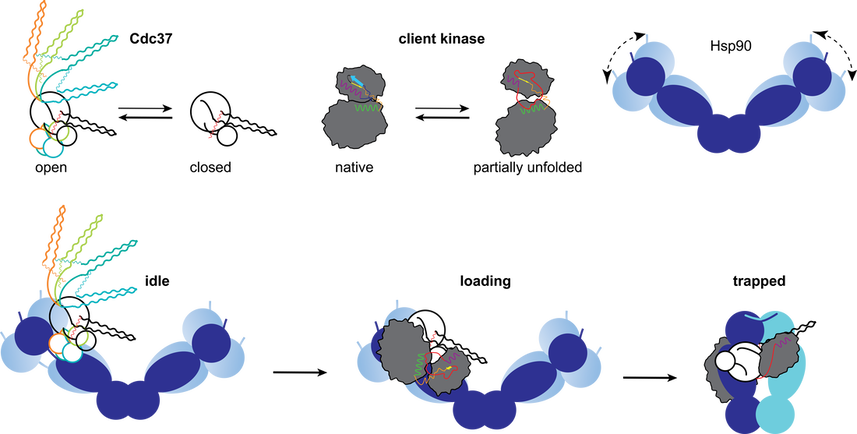
Efficient processing of Hsp90 clients involves a dynamic chaperone complex composition that is being altered in a sequential and directional manner, as well as a dynamic remodeling of its architecture, where both Hsp90-cochaperone and Hsp90/cochaperone-substrate interfaces are being reformed. At the same time protein motions (dynamics) play a fundamental role in facilitating recognition and catalysis. The modular and flexible nature of Cdc37 is instrumental to the metamorphic architecture of the Hsp90-Cdc37 complex, where distinct chaperone-cochaperone interfaces are used during substrate trapping and loading. In addition to this heterogeneous state, Cdc37 assumes a sparsely populated collapsed conformation that enables the association of client kinase domains while bound to Hsp90. Breathing motions in the free state of clients result in partial exposure of kinase motifs that become trapped by the Hsp90 dimer during inactivation. During loading, this partially unfolded client conformation is stabilized by Cdc37 independently of Hsp90, enabling loading.
Keramisanou D., Vasantha Kumar M.V., Boose N., Abzalimov R. and Gelis I.*
Assembly mechanism of early Hsp90-Cdc37-kinase complexes
Sciences Advances, 8, eabm9294 (2022)
Assembly mechanism of early Hsp90-Cdc37-kinase complexes
Sciences Advances, 8, eabm9294 (2022)
Other research directions
In collaboration with scientific friends or independently, our group engages in other research activities ranging from enzymology, antimicrobial activity and drug discovery, to supramolecular protein assemblies. Stay tuned …
Pemberton O.A., Noor R.E., Kumar M.V.V, Sanishvili R., Kemp M.T., Kearns F.L., Woodcock H.L., Gelis I.*, Chen Y*.
Mechanism of proton transfer in class A β-lactamase catalysis and inhibition by avibactam
Proc Natl Acad Sci U S A, 117, 5818-5825 (2020)
Mechanism of proton transfer in class A β-lactamase catalysis and inhibition by avibactam
Proc Natl Acad Sci U S A, 117, 5818-5825 (2020)